Adduct error means m/z is wrong when peptide result is imported from a scientific library into a Tof 2D method - WKB54773
Article number: 54773
SYMPTOMS
- When a peptide is exported from results of a peptide mapping analysis to the scientific library and then imported to a Tof 2D method, the adduct used to calculate the expected m/z is wrong
- Export a peptide ID to Scientific Library:
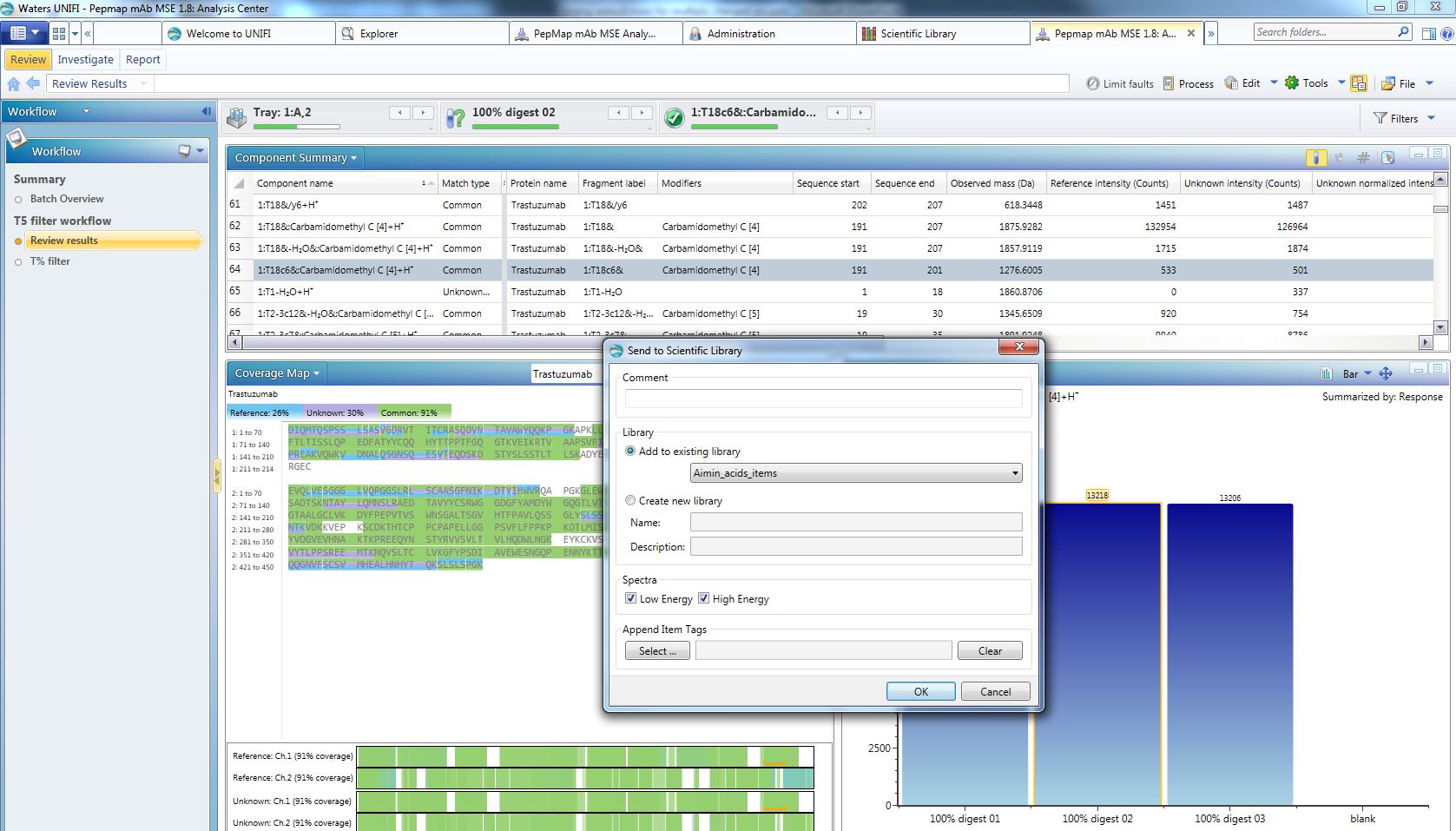
- Expected m/z shown in library:
- Create 2D Tof method:
- When the Scientific Library entry is imported into the method, the expected m/z is wrong.
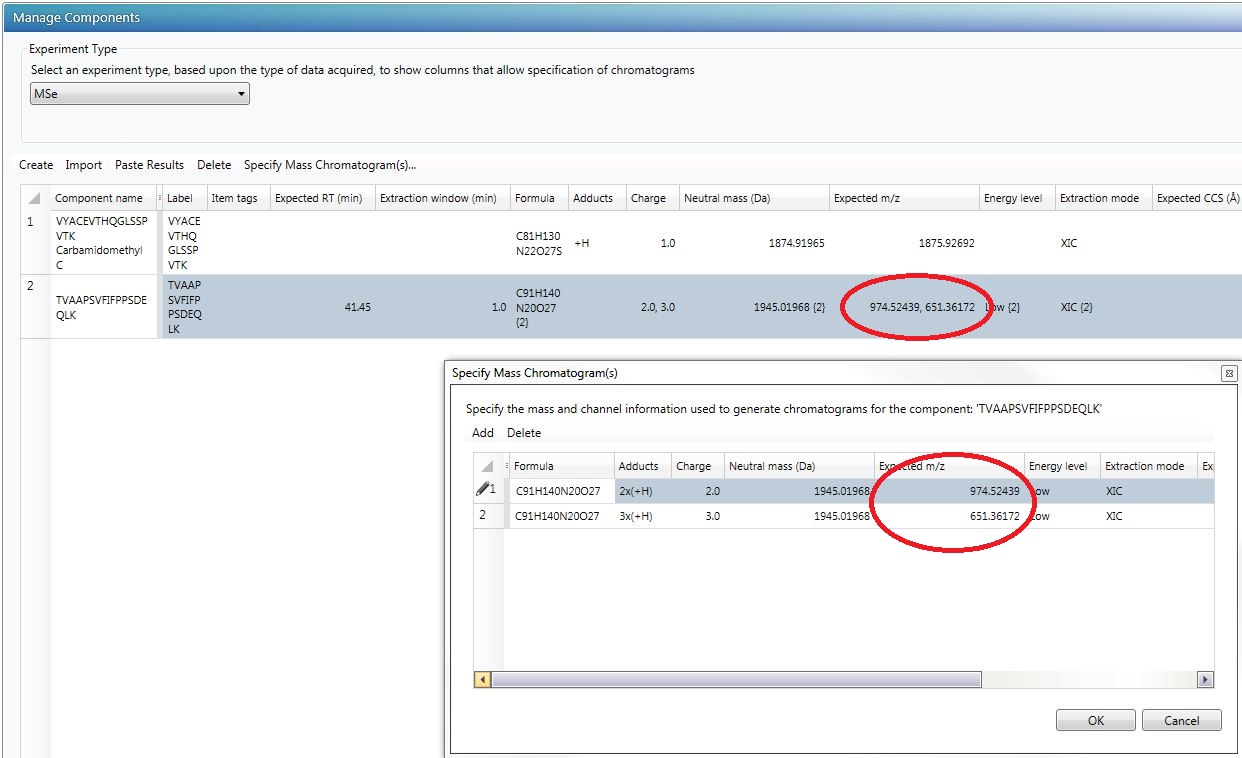
- Component masses calculated in the Manage Components section of Analysis Method > Purpose do not match the theoretical masses in the UNIFI Mass Calculator
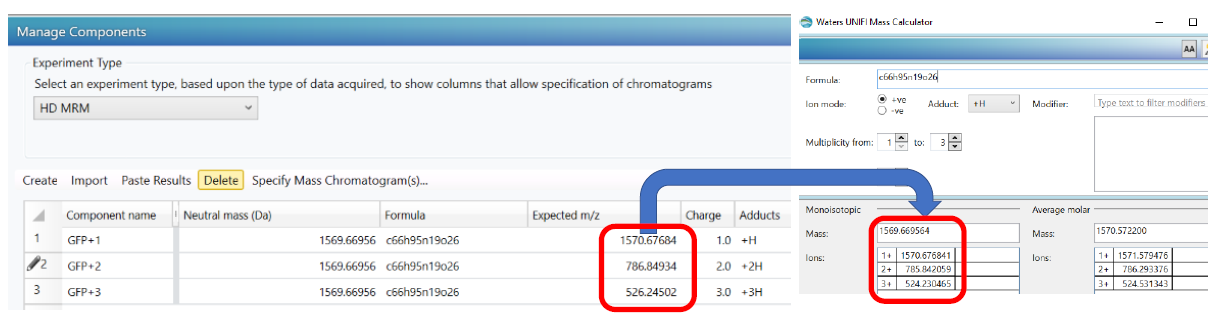
ENVIRONMENT
- UNIFI 1.9 SR4
- UNIFI 1.9 SR3
CAUSE
Bug
(It seems the H+ mass is added twice before the neutral mass is divided by the charge to generate the m/z.
FIX or WORKAROUND
Create custom 2+ and 3+ (and other) adducts and manually select them in the adducts column of the "Specific Mass Chromatogram(s)" pane. This forces the method generator to redo the m/z calculation and get it right.
ADDITIONAL INFORMATION
Raised as fault INFULA-812.
According to the fault record, this problem is fixed in UNIFI 1.9.9 and later and did not occur in UNIFI 1.8.2.

